Wellcome Genome Campus Advanced Courses and Scientific Conferences are piloting seed funding year-long research projects in Latin America and the Caribbean to extend the impact of their Genomics and Epidemiological Surveillance of Bacterial Pathogens Course, held at the University of Costa Rica, San Jose, 9-14 July 2017.
The course participants will apply the skills they learnt to real-life surveillance of bacterial pathogens across the region, identifying genomic diversity and antimicrobial resistance of Salmonella enterica, a food-borne pathogen, and Klebsiella pneumoniae, which causes pneumonia. This activity will significantly increase knowledge about the spread and emergence of these pathogens as well as their resistance to antibiotics, information which will be used to inform strategies for control and treatment. The projects will be funded by Advanced Courses and Scientific Conferences, the Centre for Genomic Pathogen Surveillance, and the Wellcome Trust Cambridge Centre for Global Health Research.
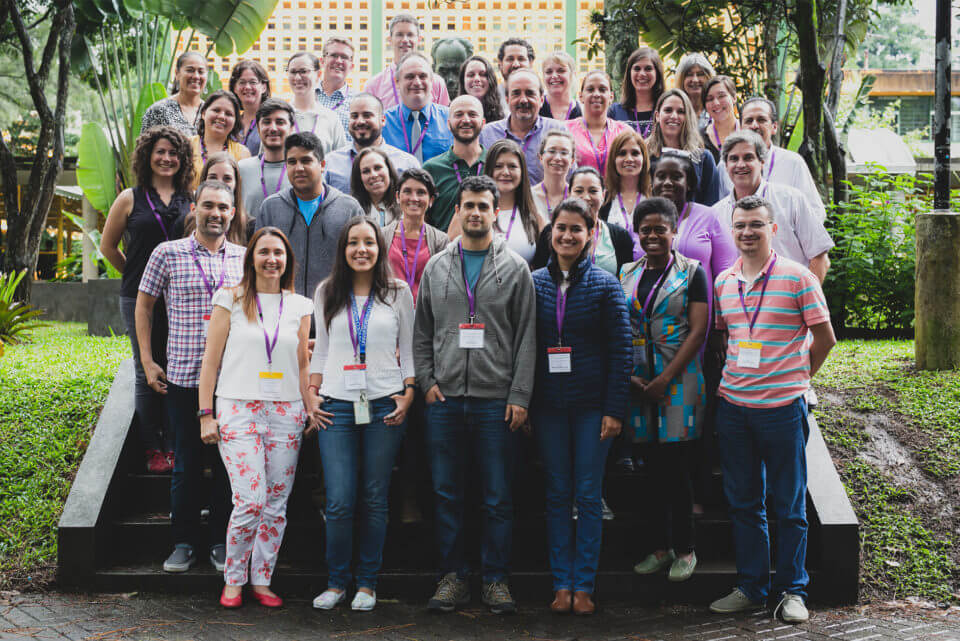

The Genomics and Epidemiological Surveillance of Bacterial Pathogens Course was established in 2013 and led by Professor Nicholas Thomson from the Wellcome Trust Sanger Institute, supported by expert instructors in genomics sciences, microbiology, and public health from a wide range of settings in Latin America, Europe and Asia, such as genomic research centres and regional and international public health organisations.
The course aims to link traditional methods of epidemiological surveillance, modern molecular typing methods, and whole genome analysis. As well as introducing these molecular techniques to these diagnostic and research laboratories, the course has resulted in strengthened networks and collaboration among course participants and instructors, both regionally and internationally.
In 2017, twenty clinical molecular biologists and microbiologists from eleven countries in Latin America and the Caribbean attended the course. This provided them with skills in sequencing techniques, anti-microbial resistance testing, phylogeny, genome assembly, and analysis through a combination of lectures, seminars, and laboratory practicals.
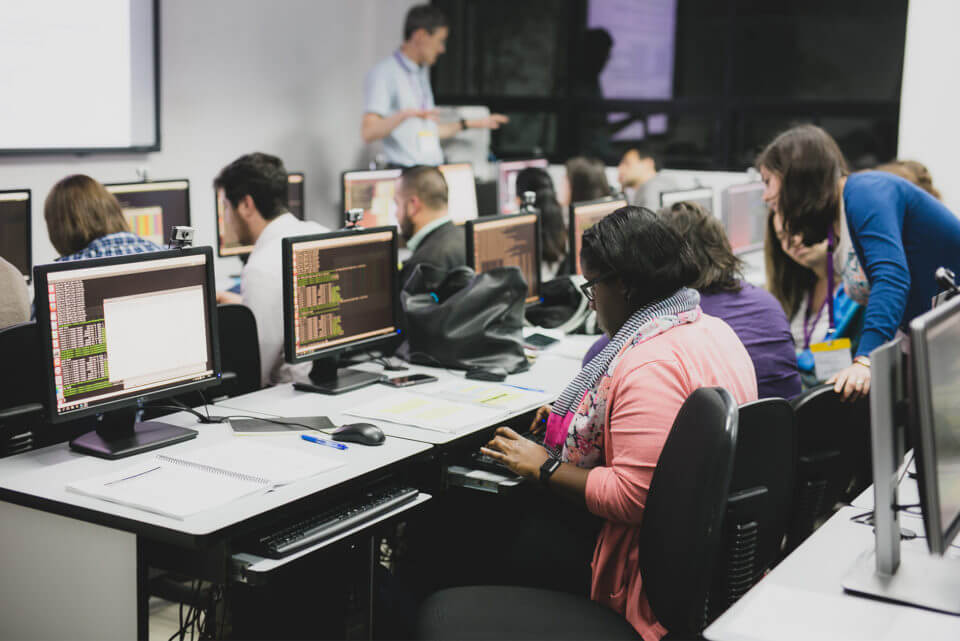
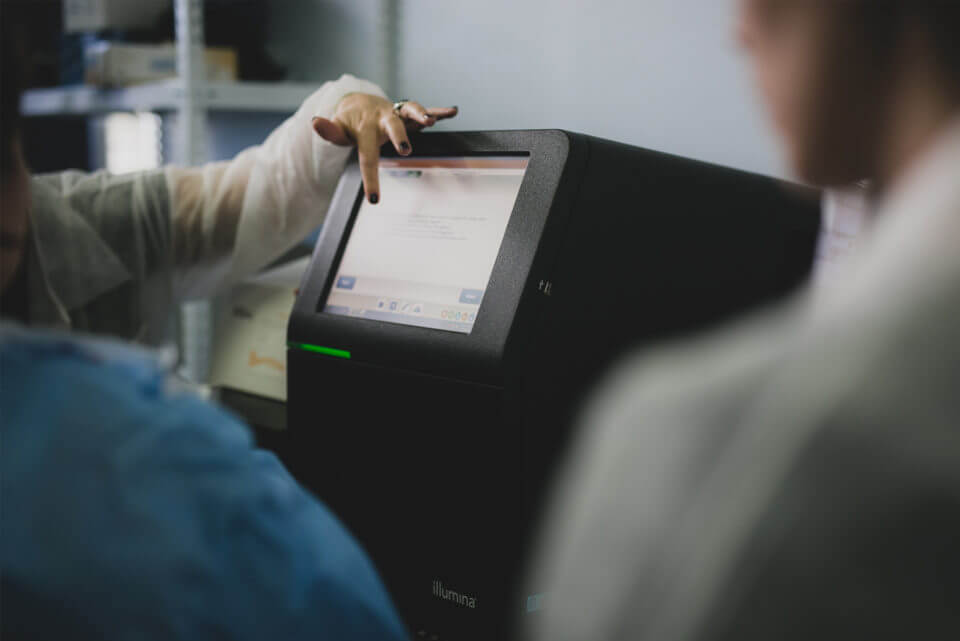




Other activities included an EpiCollect exercise demonstrated the use of smartphones in epidemiology data collection and surveillance, and a World Café discussion session, during which participants highlighted key disease topics currently important in the region, discussing key techniques and requirements to strengthen capacity for infectious disease surveillance.
This work culminated in the research projects initiative: Participants were guided through the development of collaborative research projects which aim to answer questions of public health relevance in the region through epidemiology and genomic surveillance. Having pitched their proposals to a panel of judges, funding was awarded for the implementation of two research projects: The research teams will work collaboratively to study genomic diversity and antimicrobial resistance of Salmonella enterica and Klebsiella pneumoniae by applying epidemiological methods and whole genome sequencing techniques.
This is the first time that research projects have been integrated into such a course. The participants will be expected to apply their enhanced knowledge and experience in developing regional strategies to strengthening capacity for epidemiological and genomic surveillance and anti-microbial resistance research. As well as the clear regional impact this work will have, participants will gain valuable experience in networking and implementing projects as a team through practical collaboration.
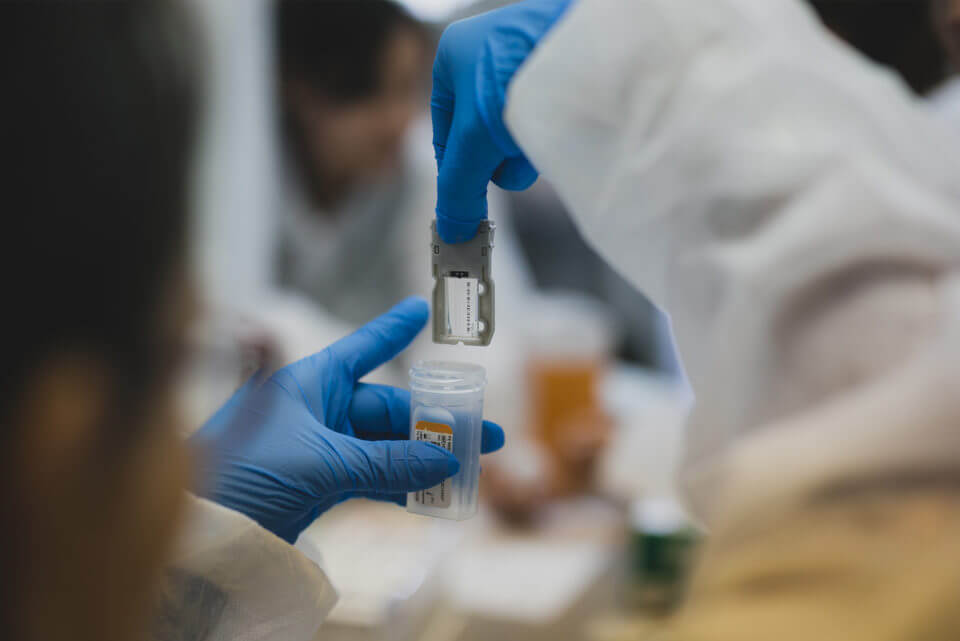
Photos by Pablo Tsukayama.